Tags
algae, biology, bloom, Ehux, Emiliania huxleyi, Genome, Popular science, science, Species, White Cliffs of Dover
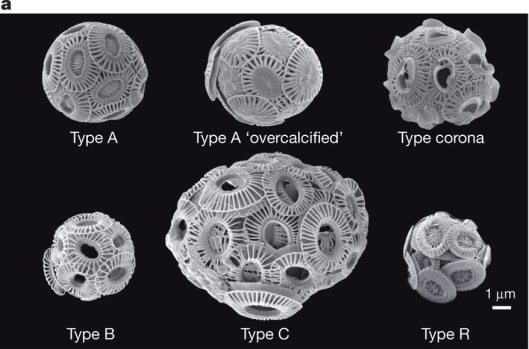
 Emiliania huxleyi has more going for it than just a beautiful name. Despite being only a few millionths of a millimeter in size — about a tenth of the thickness of a human hair — this unicellular alga has a major impact on our planet. Blooms of E. huxleyi, which can cover more than 100,000 square kilometres of ocean, are visible from space and affect the global climate; the concerted impact of all the cells in the bloom influences carbon and sulphur cycles and even changes how much light the Earth reflects. Under an electron microscope, E. huxleyi cells reveal their striking, alien beauty; encased in tiny plates called coccoliths, they look like strange spaceships or escape pods. E. huxleyi lives throughout the world’s oceans, from the warm tropics to the subarctic seas, and these plates underlie its remarkable impact on the planet’s climate and geology.
Emiliania huxleyi has more going for it than just a beautiful name. Despite being only a few millionths of a millimeter in size — about a tenth of the thickness of a human hair — this unicellular alga has a major impact on our planet. Blooms of E. huxleyi, which can cover more than 100,000 square kilometres of ocean, are visible from space and affect the global climate; the concerted impact of all the cells in the bloom influences carbon and sulphur cycles and even changes how much light the Earth reflects. Under an electron microscope, E. huxleyi cells reveal their striking, alien beauty; encased in tiny plates called coccoliths, they look like strange spaceships or escape pods. E. huxleyi lives throughout the world’s oceans, from the warm tropics to the subarctic seas, and these plates underlie its remarkable impact on the planet’s climate and geology.
The coccoliths are made of calcium carbonate — the same stuff that makes up limestone, chalk, and marble. Since the production of calcium carbonate releases CO2, E. huxleyi acts as a carbon source. It’s also photosynthetic, though, like most other algae, and so acts as a carbon sink, using sunlight to pull CO2 out of the atmosphere and convert it into food. The balance between these two dynamic processes decides whether E. huxleyi acts as a source or sink overall, leading to complex effects on the carbon cycle, particularly during massive blooms. The coccoliths also reflect light, so huge blooms also change how much heat that area of ocean absorbs.
 When E. huxleyi die, the coccoliths pull them down to the bottom of the ocean. This can be especially dramatic when a virus sweeps through a huge bloom, killing huge numbers of E. huxleyi and releasing their exoskeletons to sink. The exoskeletons pile atop one another, eventually compressing into chalk; over time, geological processes expose this rock in impressive formations like the famed White Cliffs of Dover or the Chalk Cliffs of Rügen.
When E. huxleyi die, the coccoliths pull them down to the bottom of the ocean. This can be especially dramatic when a virus sweeps through a huge bloom, killing huge numbers of E. huxleyi and releasing their exoskeletons to sink. The exoskeletons pile atop one another, eventually compressing into chalk; over time, geological processes expose this rock in impressive formations like the famed White Cliffs of Dover or the Chalk Cliffs of Rügen.
Recently, an international team of scientists finished sequencing the genome of this amazing microscopic alga. Their findings, published in a paper in Nature, offer an explanation of how E. huxleyi is able to thrive throughout the Earth’s oceans. The researchers first sequenced the genome of a particular strain of E. huxleyi. They identified around 30,000 putative genes, including many responsible for various biochemical processes, such as carbon and calcium transport and metabolism. They then used high-throughput sequencing technologies to have a look at the genomes of thirteen other strains for E. huxleyi from around the world. To their surprise, they found extensive variation between the genomes. While the different strains all share a “core genome”, each seems to supplement that core set with genes suited to its habitat, allowing E. huxleyi to cope with a wide range of environmental conditions.
“For example, if the genetic information of two humans is compared, an agreement of about 99 percent is found. However, if for example, we take two Ehux strains from different ocean regions, we find a degree of similarity of only 70 or 80 percent,” said Uwe John, one of the researchers, in a press release. “We call this phenomenon a pan-genome‘,” he continued. This is the first time a pan-genome has been found in a species of algae. The concept of a ‘pan-genome’ was developed to cope with the fluid and variable genomes of bacterial species; each pan-genome consists of a core set of genes found in all strains of a particular species as well as ‘dispensable’ genes which are only found in some of the strains.
The pan-genome of Emiliania huxleyi raises all kinds of interesting questions about what ‘species’ really means, as well as providing an excellent opportunity to study the processes of speciation and adaptation to different climate conditions. It’s also a wonderful evolutionary strategy, enabling this tiny calcified algae to spread all across the world, where it bursts forth in magnificent blooms which can reshape the environment and, eventually, form part of the coastlines of the world.
Refs
Read BA, Kegel J, Klute MJ, Kuo A, Lefebvre SC, Maumus F, Mayer C, Miller J, Monier A, Salamov A, Young J, Aguilar M, Claverie JM, Frickenhaus S, Gonzalez K, Herman EK, Lin YC, Napier J, Ogata H, Sarno AF, Shmutz J, Schroeder D, de Vargas C, Verret F, von Dassow P, Valentin K, Van de Peer Y, Wheeler G, Emiliania huxleyi Annotation Consortium, Allen AE, Bidle K, Borodovsky M, Bowler C, Brownlee C, Mark Cock J, Elias M, Gladyshev VN, Groth M, Guda C, Hadaegh A, Debora Iglesias-Rodriguez M, Jenkins J, Jones BM, Lawson T, Leese F, Lindquist E, Lobanov A, Lomsadze A, Malik SB, Marsh ME, Mackinder L, Mock T, Mueller-Roeber B, Pagarete A, Parker M, Probert I, Quesneville H, Raines C, Rensing SA, Riaño-Pachón DM, Richier S, Rokitta S, Shiraiwa Y, Soanes DM, van der Giezen M, Wahlund TM, Williams B, Wilson W, Wolfe G, Wurch LL, Dacks JB, Delwiche CF, Dyhrman ST, Glöckner G, John U, Richards T, Worden AZ, Zhang X, & Grigoriev IV (2013). Pan genome of the phytoplankton Emiliania underpins its global distribution. Nature PMID: 23760476
Medini D, Donati C, Tettelin H, Masignani V, & Rappuoli R (2005). The microbial pan-genome. Current opinion in genetics & development, 15 (6), 589-94 PMID: 16185861
The press release from the Alfred-Wegener Institute


You must be logged in to post a comment.