Tags
Agriculture, biology, breeding, Chromosome, farming, Heterosis, hybrid, hybrid vigor, Meiosis, Popular science, science, Seed
 In a really neat piece of work based around a remarkably simple bit of engineering and some textbook genetics, a team of scientists has found a way to regenerate a plant’s parents through breeding — a technique they call “reverse breeding”. This clever bit of research, which is described in a paper appearing in Nature Genetics, should be applicable to a wide range of crop species, opening up the possibility of significant advances in crop improvement and breeding programmes.
In a really neat piece of work based around a remarkably simple bit of engineering and some textbook genetics, a team of scientists has found a way to regenerate a plant’s parents through breeding — a technique they call “reverse breeding”. This clever bit of research, which is described in a paper appearing in Nature Genetics, should be applicable to a wide range of crop species, opening up the possibility of significant advances in crop improvement and breeding programmes.
Most crops today are grown from hybrid (or F1) seed, which is produced by using two highly inbred lines as parents. The resulting plants have very little genetic variation and quite uniform growth, which is important for modern, mechanized farming. The hybrids also tend to perform better and have higher yields than non-hybrids. The source of this “hybrid vigor” is poorly understood, though one possibility is that the genetic input from each parental line serves to mask the shortcomings of the other, resulting in hybrid plants that are superior to either parent. Unfortunately, hybrid vigor and uniformity is lost in subsequent generations because of the mixture of genetic material during sex. Breeders therefore have to continuously re-create the hybrids by crossing the inbred parental lines. This means that any improvements have to be bred into the parental lines; it also restricts the breeders’ ability to take advantage of outbreeding in their breeding programme.
The loss of hybrid vigor is because of genetic variation introduced during meiosis, the production of eggs and sperm. Most of this variation isn’t from mutation but is the result of two processes called assortment and crossing over. Our DNA is packaged into chromosomes which are arranged in pairs, each of which consists of one maternal and one paternal chromosome. Assortment simply means that each pair is randomly divided during meiosis, resulting in each sperm or egg cell having a random mixture of maternal and paternal chromosomes. In addition, before this division happens the chromosomes of each pair “cross over”; they swap bits of DNA between themselves and so are no longer intact maternal or paternal chromosomes, but rather a mixture of DNA from both.
Using playing cards as an analogy, everyone starts with a hand made up of pairs of red and black cards, representing maternal and paternal chromosomes. During meiosis, only half of the hand goes into each sperm or egg cell; assortment is simply shuffling before passing along cards during meiosis — there’s no guarantee that the cards chosen will be the same colour, but the individual cards are all still intact. By contrast, crossing over would mean cutting out parts of one card from a pair and gluing them into the other before shuffling. The resulting cards would be a mish-mash of red and black, making it impossible to recreate the original hand. If this step could be avoided, then recovering the original (parental) hand would just be a question of reshuffling enough times.
That’s precisely what the researchers did. Since crossing over scrambles chromosomes into a mix of maternal and paternal DNA, it becomes impossible to regenerate the original parents that were bred to create the hybrid. By blocking crossing over, the researchers created plants which passed along their maternal and paternal chromosomes intact, making meiosis nothing more than a reshuffling of chromosomes. Each sperm or egg cell therefore contained a random combination of intact maternal and paternal chromosomes; reverse breeding to regenerate the original parental lines is then simply a matter of juggling chromosomes. This task was made much easier by the fact that it’s possible to recreate an entire plant from a single sperm or egg; there are various techniques for doing this, but the basic idea is to double the DNA, making each chromosome duplicate itself to form a pair. By generated plants from sperm and eggs which hadn’t undergone crossing over, the researchers were able to start with a hybrid plant where every pair of chromosomes 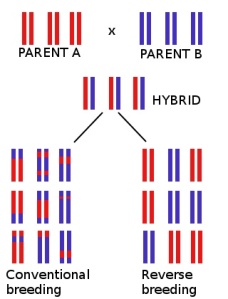 was composed of one chromosome from each parent and breed plants in which each pair of chromosomes was made of two identical copies from the same parent. Recovering the original parents was then simply a matter of crossing these plants in the right combination. In fact, this technique doesn’t just recreate the original parents; it also recreates several sets of possible parents — that is, pairs of plants which, if bred together, would produce the hybrid. For example, the second and third reverse bred plants in the figure aren’t the original parents, but a cross between them would also generate the hybrid.
was composed of one chromosome from each parent and breed plants in which each pair of chromosomes was made of two identical copies from the same parent. Recovering the original parents was then simply a matter of crossing these plants in the right combination. In fact, this technique doesn’t just recreate the original parents; it also recreates several sets of possible parents — that is, pairs of plants which, if bred together, would produce the hybrid. For example, the second and third reverse bred plants in the figure aren’t the original parents, but a cross between them would also generate the hybrid.
In order to block crossing over, the researchers reduced the expression of a gene called DMC1 which produces an enzyme that is important for this process. Since DMC1 is widely conserved between different plant species, it should be straightforward to apply this technique in different crops even though this study used Arabidopsis thaliana, a small relative of mustard which serves as the model organism for plant genetics. By making sure that the DMC1 reduction was dominant, the researchers were able to be sure that the regenerated plants weren’t harboring an unseen copy; this is quite important since plants with decreased DMC1 also had lower fertility, which would be unacceptable in crops.
It’s clear that this work will have important practical applications for plant breeders. It also makes it quite straightforward to create chromosome-substitution lines — that is, plants which have one chromosome from variety A and the rest from variety B; this will be a useful tool for breeders and for plant biologists in general. My favourite thing about this study, though, isn’t the practical benefits it will generate but rather its striking simplicity; like a lot of the best scientific work, it seems so obvious and straightforward in retrospect that I wish I’d been clever enough to think of it.
Ref:
Wijnker, E., van Dun, K., de Snoo, C., Lelivelt, C., Keurentjes, J., Naharudin, N., Ravi, M., Chan, S., de Jong, H., & Dirks, R. (2012). Reverse breeding in Arabidopsis thaliana generates homozygous parental lines from a heterozygous plant Nature Genetics, 44 (4), 467-470 DOI: 10.1038/ng.2203

An Approach to the Genetic Improvement of Clonal Cultivars via Backcrossing
Wolfgang Link and Albrecht E. Melchinger
Vol. 35 No. 3, p. 931-931
Published: May, 1995
Maybe you, as I do, ask yourself (after reading the short note cited above) why the authors of this inspiring bit of work no not cite this ‘research topic letter’ – in spite of being well informed about its existance. I mean, what does it cost them to admit earlier ideas?
Unfortunately I can’t access your paper, so I can’t really comment on this, but I’m sorry to hear that you weren’t cited when you think you should have been! Science is a collaborative effort in which acknowledging the contributions of others plays a central role. Of course, mistakes get made (I’m sure I’m guilty of some myself), but you seem to think this was an intentional omission. I think it’s great that you’ve raised the issues so everyone can be aware of it and have a look at your paper, though it’s a shame many people won’t be able to access it.
This old, short note can be accessed via http://www.uni-goettingen.de/en/48273.html; you find it there as ‘Research Topic Letter’.
Due to our current discussion, I re-read the paper of Wijnker et al., in Nature Genetics (44;4) 2012. Since this topic deals with Arabidopsis, there may be still another application than those mentioned there. They rightly say that many-chromosome crops such as canola (which is a relative of A.t.) are not quite fit for reverse-breeding mutually complementing inbreds from a hybrid. Well. What still could be done in canola, based on these ideas, is a ‘decomposition’ of the allotetraploid genome of Brassica napus into the two diploid genomes of its ancestral (crop) species, B. rapa (turnip rape) and B. olearacea (cabbage). Some of the anouploid reverse-bred dihaploids will – by chance – consist of just the chromosomes of the A genome and/or of the C genome. These need not be many, and with x=19 chromosomes these will for sure not be many. Yet there is no need to have many for the invisaged purpose. The purpose would be to ‘phenotypically see’ which type of B. rapa and which type of caggabe is ‘living’ in our current-day canoly – with presumably many consequences for further canola breeding.
Probably somebody already proposed this or is even currently doing this. Fine! If not, my comment dates from today (May 21 2013). I participated in a discussion of such decomposition already several years ago, but with the reverse-breeding idea and the corresponding genes (DMC1, CENH3) this should now – thanks to Wijnker et al. – be ready-to-do it.
Dear Dr. Link, I understand your dissatisfaction of not getting cited in Wijnker’s paper. It’s OK. However, I would like to make some points. As it reads, your research note mostly deals with genetic improvement of clonal cultivars through backcrossing. Your paper is quite different from this paper in three aspects; Wijnker et al have attempted targeted engineering (and not the routine straightforward bulky plant breeding), have addressed a seed propagated cross pollinated plant (not clonally propagated plant) using “one step” approach (not by a series of backcrossings). Therefore it acquires novelty in its own merit.
Secondly, you have discussed about “decomposition” of the parental genomes in B. napus to their respective lineages (AACC into AA and CC, if I have understood you properly). This may be possible in crops like Brassica where genome evolution is known (as in the form of U triangle) and breeders have undertaken resynthesis of tetraploid Brassicas from their progenitor diploid parents previously. Obtaining present day commercially useful tetraploids by resynthesis is a herculian task, if not impossible. Also, this method simply does not work for other crops where genome evolution is not so straightforward as in Brassicas.
I may have erred in my points; however, largely, this is what I believe that Wijnker and Ravi’s labs’ research are demonstrably fresh with a lot of future commercial stakes (benefitting farmers!).
Dear Wolfgang, Mon 10/03/2014 12:57
you might have come across it already, but please find attached our most recent publication on reverse breeding.
As you will see, we now made reference to your 1995 article in Crop Science, which, as you previously suggested, does merit attention in relation to reverse breeding.
With my best regards,
Erik Wijnker
Published online 6 March 2014; doi:10.1038/nprot.2014.049; Nature America Inc.;
Hybrid recreation by reverse breeding in Arabidopsis thaliana
Erik Wijnker et al., 2014
Thank you very much, dear Erik, Mon 10/03/2014 13:49
for this nice info, I feel very much pleased (and, I admit, satisfied).
With best greetings from Göttingen,
Wolfgang Link
PS I will of course carefully study your current paper.
🙂 🙂 🙂